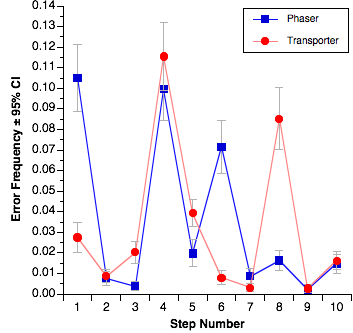
Residues A2058 and A2059 of 23S rRNA form a hydrophobic crevice into which macrolide antibiotics bind. All ribosomal proteins except L4 and L22 have been omitted from the rendering. coli large ribosome subunit viewed from the cytoplasmic opening of the exit tunnel. L4 and L22 loops and the ribosomal macrolide-binding site. Although less common than Erm-mediated resistance, L4 and L22 mutations are increasingly being identified in clinically relevant species such as Streptococcus pneumoniae, Haemophilus influenzae, and Staphylococcus aureus 13 14 15. L4 and L22 each have extended loops, which converge to form a narrowing in the exit tunnel adjacent to the macrolide-binding site ( Fig. Finally, the macrolide-binding site can be modified by mutations in rplD and rplV, which encode ribosomal proteins L4 and L22, respectively. Mutation of A2058, A2059, and other 23S rRNA residues within the macrolide-binding site can confer a similar resistance phenotype, but these mutations are typically limited to bacteria containing only one or two rRNA operons 10. 1b), and its methylation confers resistance to all three classes of antibiotics 8 9. A2058 is critical for the binding of macrolides, lincosamides, and streptogramin B-type antibiotics ( Fig. 
Modification of the macrolide-binding site is the most widespread resistance mechanism and is usually associated with the acquisition of Erm methyltransferases, which catalyze N 6, N 6-dimethylation of A2058 in 23S rRNA ( Escherichia coli numbering is used throughout) 8 11 12. Resistance to macrolides is mediated by three general mechanisms: i) enzymatic inactivation of macrolides, ii) increased macrolide efflux, and iii) alteration of the macrolide-binding site 8 9 10. In addition, macrolides have been shown to interfere with ribosome assembly in several bacterial species 5 6 7. These drugs bind to the large ribosomal subunit at the entrance of the nascent peptide exit tunnel, near the peptidyl transferase center, where they inhibit peptide bond formation and facilitate peptidyl-tRNA dissociation from the ribosome 1 2 3 4. Macrolide antibiotics – which include erythromycin and tylosin – are commonly used antibacterial agents that interfere with protein synthesis. This work underscores the exceptional functional plasticity of the L4 and L22 proteins, and highlights the utility of Red-mediated recombination in targeted genetic selections. Moreover, DMS methylation protection assays demonstrated that L22 Lys90Trp ribosomes bind tylosin more readily than erythromycin in vivo. Purified L22 Lys90Trp ribosomes show reduced erythromycin binding, but have the same affinity for tylosin as wild-type ribosomes. L22 Lys90Trp is one such allele, which confers resistance to erythromycin, but not tylosin or spiramycin. Although L4 and L22 mutants are typically resistant to most macrolides, selections carried out on different antibiotics revealed macrolide-specific resistance mutations. Transfer of L4 and L22 mutations into wild-type cells by phage P1-mediated transduction demonstrated that each allele was sufficient to confer macrolide resistance. Many resistance mutations were complex, involving multiple missense mutations, in-frame deletions, and insertions. These experiments led to the identification of 341 different resistance mutations encoding 278 unique L4 and L22 proteins – the overwhelming majority of which are novel. We randomized residues at the tips of the L4 and L22 loops using recombineered oligonucleotide libraries, and selected the mutagenized cells for erythromycin-resistant mutants. Here, we use bacteriophage λ Red-mediated recombination, or “recombineering”, to uncover new L4 and L22 alleles that confer macrolide resistance in Escherichia coli. L4 and L22 have elongated loops whose tips converge in the peptide exit tunnel near the macrolide binding site, and resistance mutations typically affect residues within these loops. Mutations in ribosomal proteins L4 and L22 confer resistance to erythromycin and other macrolide antibiotics in a variety of bacteria.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
